Helixyte™ Green Fluorimetric dsDNA Quantitation Kit
Optimized for Broad Dynamic Range
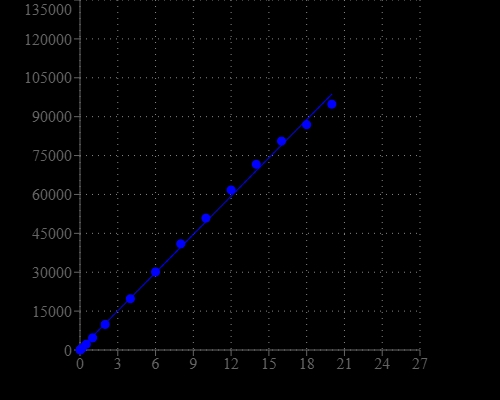
DNA Quantitation is a very important task in DNA sample preparations for various analyses. Helixyte™ Green Fluorimetric dsDNA Quantitation Kit provides a rapid method to quantify dsDNA with Helixyte™ Green BR . The assay is linear over three orders of magnitude and is a few magnitudes more sensitive than UV absorbance readings. Helixyte™ Green BR exhibits large fluorescence enhancement upon binding to dsDNA and has little sequence dependence, allowing to the accurate measurement of DNA samples from various sources, including genomic DNA, viral DNA, miniprep DNA or PCR amplification products. The assay is highly selective for double-stranded DNA (dsDNA) over RNA and is optimized to measure DNA concentrations from 10 pg/µL to 10 ng/µL.

| Catalog | Size | Price | Quantity |
|---|---|---|---|
| 17645 | 200 Tests | Price | |
| 17646 | 1000 Tests | Price |
Spectral properties
| Excitation (nm) | 503 |
| Emission (nm) | 527 |
Storage, safety and handling
| Intended use | Research Use Only (RUO) |
| Storage | Freeze (< -15 °C); Minimize light exposure |
Instrument settings
| Fluorescence microplate reader | |
| Excitation | 490 nm |
| Emission | 530 nm |
| Cutoff | 515 nm |
| Recommended plate | Solid black |
Contact us
| Telephone | |
| Fax | |
| sales@aatbio.com | |
| International | See distributors |
| Bulk request | Inquire |
| Custom size | Inquire |
| Technical Support | Contact us |
| Request quotation | Request |
| Purchase order | Send to sales@aatbio.com |
| Shipping | Standard overnight for United States, inquire for international |
Page updated on May 8, 2026

