ATP-TAMRA conjugate [5-TAMRA-eda-ATP]
Ordering information
| Price | |
| Catalog Number | |
| Unit Size | |
| Quantity |
Additional ordering information
| Telephone | 1-800-990-8053 |
| Fax | 1-800-609-2943 |
| sales@aatbio.com | |
| International | See distributors |
| Bulk request | Inquire |
| Custom size | Inquire |
| Shipping | Standard overnight for United States, inquire for international |
Physical properties
| Molecular weight | 1128.13 |
| Solvent | DMSO |
Spectral properties
| Correction Factor (260 nm) | 0.32 |
| Correction Factor (280 nm) | 0.178 |
| Extinction coefficient (cm -1 M -1) | 90000 |
| Excitation (nm) | 552 |
| Emission (nm) | 578 |
Storage, safety and handling
| H-phrase | H303, H313, H333 |
| Hazard symbol | XN |
| Intended use | Research Use Only (RUO) |
| R-phrase | R20, R21, R22 |
| Storage | Freeze (< -15 °C); Minimize light exposure |
| UNSPSC | 12352200 |
| Overview |
Molecular weight 1128.13 | Correction Factor (260 nm) 0.32 | Correction Factor (280 nm) 0.178 | Extinction coefficient (cm -1 M -1) 90000 | Excitation (nm) 552 | Emission (nm) 578 |
Fluorescently labeled ATP tracers are useful for monitoring ATP-binding proteins and other biomolecules. The ATP-TAMRA tracer was demonstrated to bind ATPases that are a class of enzymes that catalyze the decomposition of ATP into ADP and a free phosphate ion.
Calculators
Common stock solution preparation
Table 1. Volume of DMSO needed to reconstitute specific mass of ATP-TAMRA conjugate [5-TAMRA-eda-ATP] to given concentration. Note that volume is only for preparing stock solution. Refer to sample experimental protocol for appropriate experimental/physiological buffers.
| 0.1 mg | 0.5 mg | 1 mg | 5 mg | 10 mg | |
| 1 mM | 88.642 µL | 443.211 µL | 886.423 µL | 4.432 mL | 8.864 mL |
| 5 mM | 17.728 µL | 88.642 µL | 177.285 µL | 886.423 µL | 1.773 mL |
| 10 mM | 8.864 µL | 44.321 µL | 88.642 µL | 443.211 µL | 886.423 µL |
Molarity calculator
Enter any two values (mass, volume, concentration) to calculate the third.
| Mass (Calculate) | Molecular weight | Volume (Calculate) | Concentration (Calculate) | Moles | ||||
| / | = | x | = |
Spectrum
Open in Advanced Spectrum Viewer
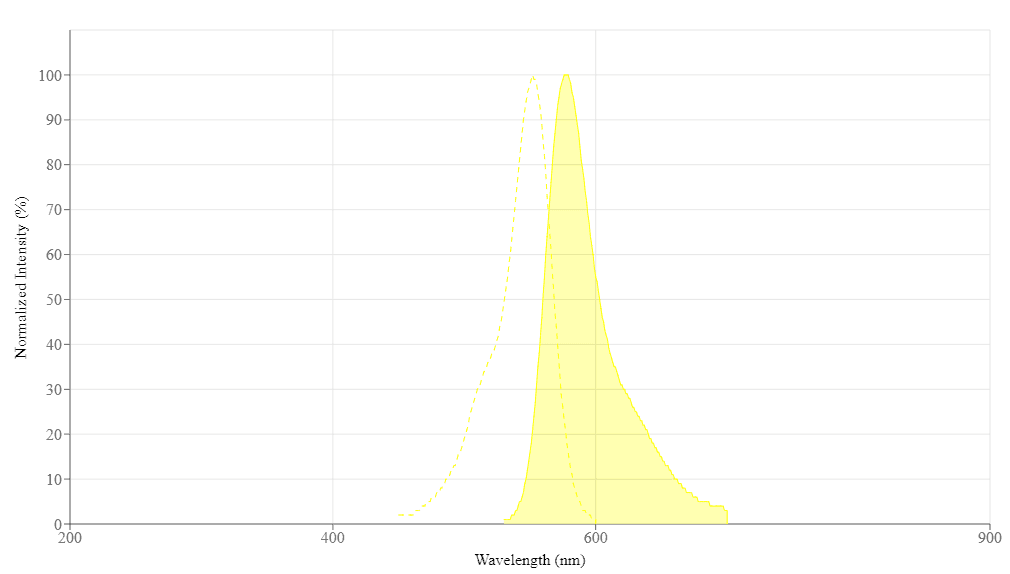

Spectral properties
| Correction Factor (260 nm) | 0.32 |
| Correction Factor (280 nm) | 0.178 |
| Extinction coefficient (cm -1 M -1) | 90000 |
| Excitation (nm) | 552 |
| Emission (nm) | 578 |
Product Family
| Name | Excitation (nm) | Emission (nm) | Extinction coefficient (cm -1 M -1) | Correction Factor (260 nm) | Correction Factor (280 nm) |
| ADP-TAMRA conjugate [5-TAMRA-eda-ADP] | 552 | 578 | 90000 | 0.32 | 0.178 |
References
View all 32 references: Citation Explorer
Deciphering the catalysis-associated conformational changes of human adenylate kinase 1 with single-molecule spectroscopy
Authors: Lin CY, Huang JY, Lo LW.
Journal: J Phys Chem B (2013): 13947
Authors: Lin CY, Huang JY, Lo LW.
Journal: J Phys Chem B (2013): 13947
The Escherichia coli PriA helicase-double-stranded DNA complex: location of the strong DNA-binding subsite on the helicase domain of the protein and the affinity control by the two nucleotide-binding sites of the enzyme
Authors: Szymanski MR, Jezewska MJ, Bujalowski W.
Journal: J Mol Biol (2010): 344
Authors: Szymanski MR, Jezewska MJ, Bujalowski W.
Journal: J Mol Biol (2010): 344
The effect of NBD-Cl in nucleotide-binding of the major subunit alpha and B of the motor proteins F1FO ATP synthase and A1AO ATP synthase
Authors: Hunke C, Tadwal VS, Manimekalai MS, Roessle M, Gruber G.
Journal: J Bioenerg Biomembr (2010): 1
Authors: Hunke C, Tadwal VS, Manimekalai MS, Roessle M, Gruber G.
Journal: J Bioenerg Biomembr (2010): 1
ATP/ADP binding to a novel nucleotide binding domain of the reticulocyte-binding protein Py235 of Plasmodium yoelii
Authors: Ramalingam JK, Hunke C, Gao X, Gruber G, Preiser PR.
Journal: J Biol Chem (2008): 36386
Authors: Ramalingam JK, Hunke C, Gao X, Gruber G, Preiser PR.
Journal: J Biol Chem (2008): 36386
Reversal of ADP-mediated aggregation of adenosine kinase by cyclophilin leads to its reactivation
Authors: Sen B, Chakraborty A, Datta R, Bhattacharyya D, Datta AK.
Journal: Biochemistry (2006): 263
Authors: Sen B, Chakraborty A, Datta R, Bhattacharyya D, Datta AK.
Journal: Biochemistry (2006): 263
ATPase mechanism of Eg5 in the absence of microtubules: insight into microtubule activation and allosteric inhibition by monastrol
Authors: Cochran JC, Gilbert SP.
Journal: Biochemistry (2005): 16633
Authors: Cochran JC, Gilbert SP.
Journal: Biochemistry (2005): 16633
Ca2+ binding to sarcoplasmic reticulum ATPase phosphorylated by Pi reveals four thapsigargin-sensitive Ca2+ sites in the presence of ADP
Authors: Vieyra A, Mintz E, Lowe J, Guillain F.
Journal: Biochim Biophys Acta (2004): 103
Authors: Vieyra A, Mintz E, Lowe J, Guillain F.
Journal: Biochim Biophys Acta (2004): 103
Evidence for proximal cysteine and lysine residues at or near the active site of arginine kinase of Stichopus japonicus
Authors: Guo Q, Chen B, Wang X.
Journal: Biochemistry (Mosc) (2004): 1336
Authors: Guo Q, Chen B, Wang X.
Journal: Biochemistry (Mosc) (2004): 1336
D1 ring is stable and nucleotide-independent, whereas D2 ring undergoes major conformational changes during the ATPase cycle of p97-VCP
Authors: Wang Q, Song C, Yang X, Li CC.
Journal: J Biol Chem (2003): 32784
Authors: Wang Q, Song C, Yang X, Li CC.
Journal: J Biol Chem (2003): 32784
Conformational dynamics of DnaB helicase upon DNA and nucleotide binding: analysis by intrinsic tryptophan fluorescence quenching
Authors: Flowers S, Biswas EE, Biswas SB.
Journal: Biochemistry (2003): 1910
Authors: Flowers S, Biswas EE, Biswas SB.
Journal: Biochemistry (2003): 1910
