Amplite® Rapid Colorimetric Maleimide Quantitation Kit
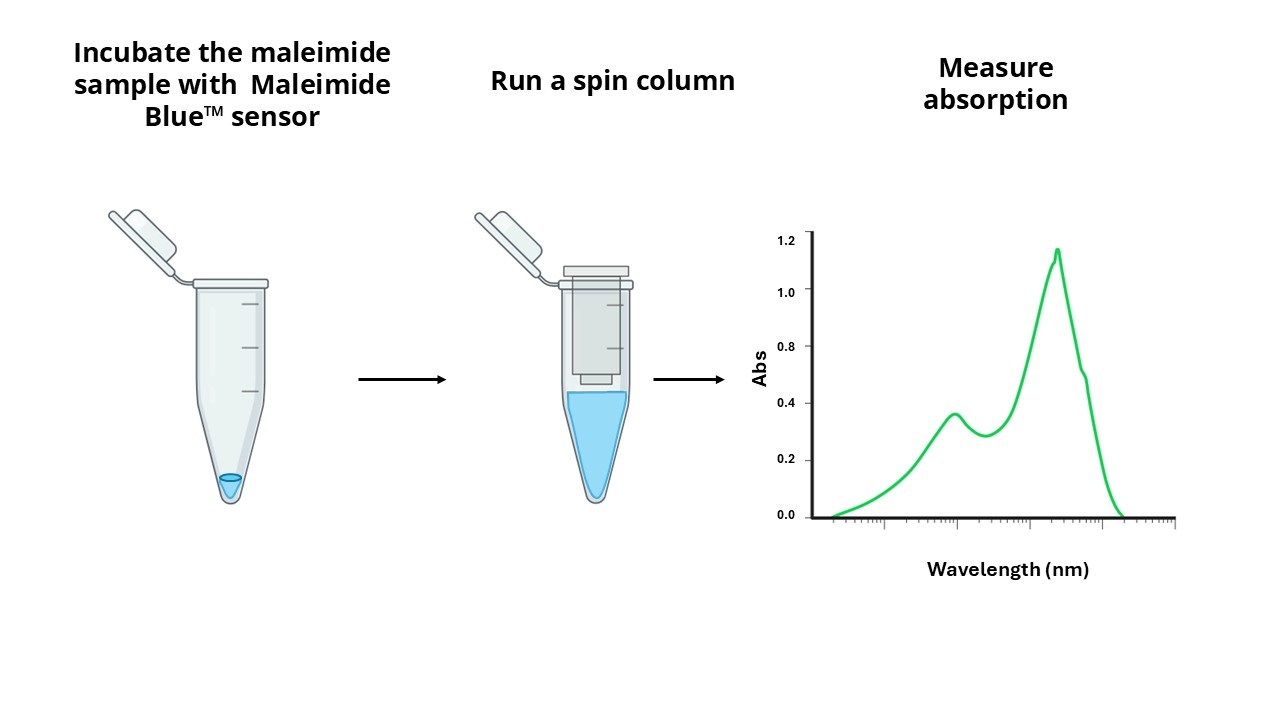
A variety of crosslinking reagents with a maleimide group are widely used for crosslinking proteins to proteins, or proteins to other biomolecules. A key challenge in maleimide crosslinking technology has been the quantitation of maleimide linked to a single protein or antibody. Maleimides can be directly assayed spectrophotometrically at 302 nm. However, the small extinction coefficient of 620 M-1cm-1 renders this assay insensitive, and the assay is further complicated by the protein absorbance at the same wavelength. AAT Bioquest's Amplite® Rapid Colorimetric Maleimide Quantitation Kit provides a rapid and accurate method to quantify maleimide using our proprietary maleimide sensor Maleimide Blue™ with the maximum absorbance at ~780nm. The principle of this assay is that Maleimide Blue™ reacts with the maleimide-linked sample, and the resulted product is run through a single spin column to remove the excess sensor. The absorption spectrum of the purified product is measured, and the amount of maleimide to protein ratio can be calculated from the absorbance ratio of 780 nm and 280 nm (for proteins) or 260 nm (for oligos and nucleic acids). This Amplite® Rapid Maleimide Quantitation kit can be performed in a traditional cuvette, NanoDrop™ Spectrophotometer or a convenient 96-well absorbance plate reader with a UV-transparent plate. This kit has been widely used for the rapid quantification of maleimide group from protein, oligo and nucleic acid samples.

| Catalog | Size | Price | Quantity |
|---|---|---|---|
| 5526 | 2 Tests | Price |
Storage, safety and handling
| H-phrase | H303, H313, H333 |
| Hazard symbol | XN |
| Intended use | Research Use Only (RUO) |
| R-phrase | R20, R21, R22 |
| UNSPSC | 12352200 |
Instrument settings
| Absorbance microplate reader | |
| Absorbance | 900 nm to 250 nm |
| Recommended plate | Clear bottom |
Contact us
| Telephone | |
| Fax | |
| sales@aatbio.com | |
| International | See distributors |
| Bulk request | Inquire |
| Custom size | Inquire |
| Technical Support | Contact us |
| Request quotation | Request |
| Purchase order | Send to sales@aatbio.com |
| Shipping | Standard overnight for United States, inquire for international |
Page updated on May 14, 2026
