MitoLite™ Red CMXRos
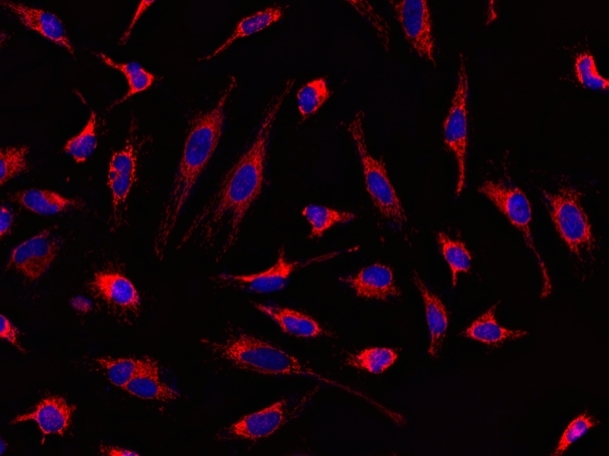
MitoLite™ Red CMXRos is chemically same to the MitoTracker™ Red CMXRos (ThermoFisher). MitoLite™ Red CMXRos is a cationic dye that selectively accumulates in mitochondria probably vial the mitochondrial membrane potential gradient. The mitochondrial indicator is a hydrophobic compound that easily permeates intact live cells, and trapped in mitochondria after it gets into cells. This fluorescent mitochondrial indicator is retained in mitochondria for long time since the indicator carries a cell-retaining group. This key feature significantly increases its staining efficiency. MitoLite™ Red CMXRos is well-retained after aldehyde fixation.

| Catalog | Size | Price | Quantity |
|---|---|---|---|
| 22698 | 10x50 ug | Price |
Physical properties
| Molecular weight | 531.52 |
| Solvent | DMSO |
Spectral properties
| Excitation (nm) | 578 |
| Emission (nm) | 598 |
Storage, safety and handling
| H-phrase | H303, H313, H333 |
| Hazard symbol | XN |
| Intended use | Research Use Only (RUO) |
| R-phrase | R20, R21, R22 |
| Storage | Freeze (< -15 °C); Minimize light exposure |
| UNSPSC | 12352200 |
| CAS | 167095-09-2 |
Instrument settings
| Fluorescence microscope | |
| Excitation | TRITC filter set |
| Emission | TRITC filter set |
| Recommended plate | Black wall/clear bottom |
Contact us
| Telephone | |
| Fax | |
| sales@aatbio.com | |
| International | See distributors |
| Bulk request | Inquire |
| Custom size | Inquire |
| Technical Support | Contact us |
| Request quotation | Request |
| Purchase order | Send to sales@aatbio.com |
| Shipping | Standard overnight for United States, inquire for international |
Page updated on June 6, 2026

