PhosphoWorks™ Colorimetric MESG Phosphate Assay Kit
UV absorption
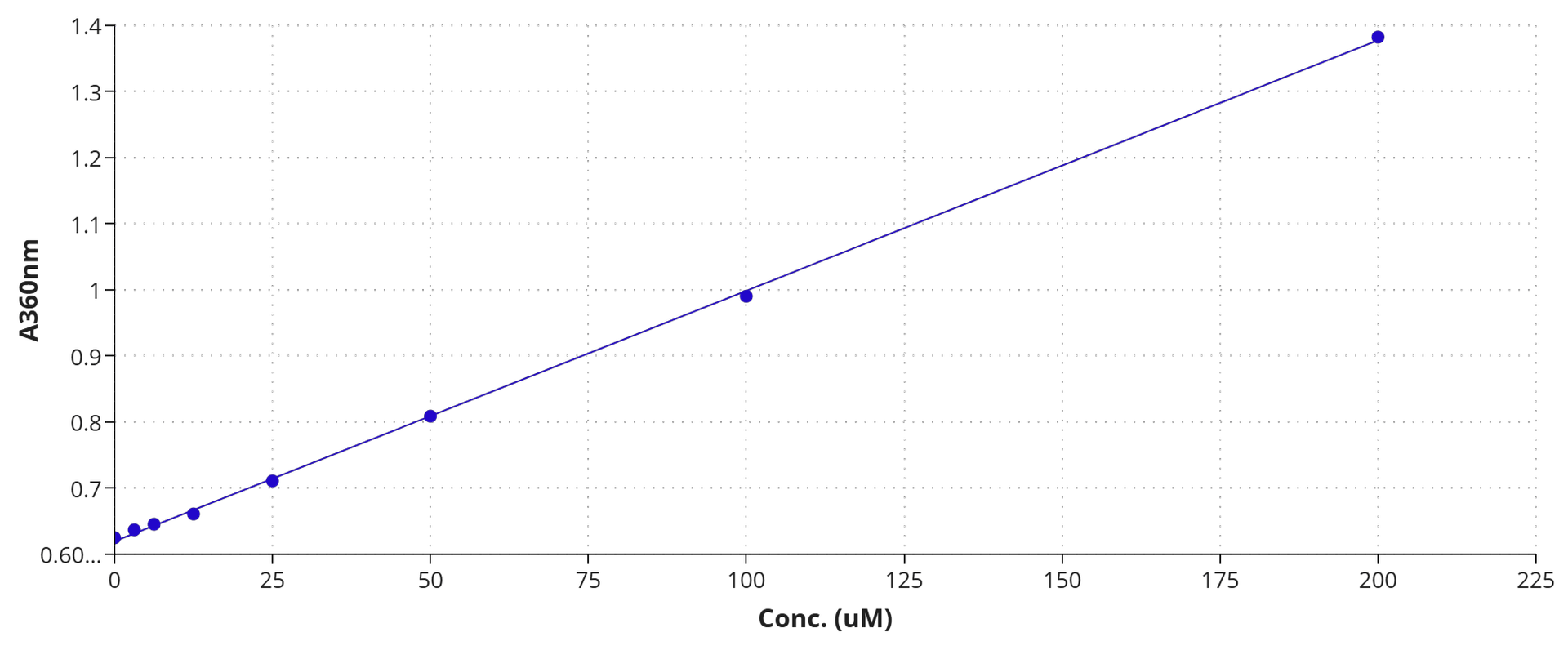
In the presence of inorganic phosphate MESG is converted to 2-amino-6-mercapto-7-methlpurine by purine nucleoside phosphorylase (EC 2.4.2.1) with absorption wavelength shift to red. This feature has been used to develop our convenient MESG phosphate assay kit. Our kit provides all the essential reagents including MESG, phosphorylase and reaction buffer. The MESG substrate gives an absorbance increase at 360 nm on phosphorylysis at pH 6.5-8.5, and at pH 7.6 the change in extinction coefficient is 11,000 M-1cm-1. The assay is shown to quantitate phosphate in solution at concentrations at least down to 2 µM. It can be used to measure the kinetics of phosphate release from phosphatases (such as GTPases and ATPases) by coupling the two enzymatic reactions.

| Catalog | Size | Price | Quantity |
|---|---|---|---|
| 21659 | 200 Tests | Price |
Storage, safety and handling
| H-phrase | H303, H313, H333 |
| Hazard symbol | XN |
| Intended use | Research Use Only (RUO) |
| R-phrase | R20, R21, R22 |
| UNSPSC | 12352200 |
Instrument settings
| Spectrophotometer | |
| Absorbance | 360 nm |
| Recommended plate | Clear UV-transparent |
| Absorbance microplate reader | |
| Absorbance | 360 nm |
| Recommended plate | Clear bottom |
Contact us
| Telephone | |
| Fax | |
| sales@aatbio.com | |
| International | See distributors |
| Bulk request | Inquire |
| Custom size | Inquire |
| Technical Support | Contact us |
| Request quotation | Request |
| Purchase order | Send to sales@aatbio.com |
| Shipping | Standard overnight for United States, inquire for international |
Page updated on June 5, 2026
