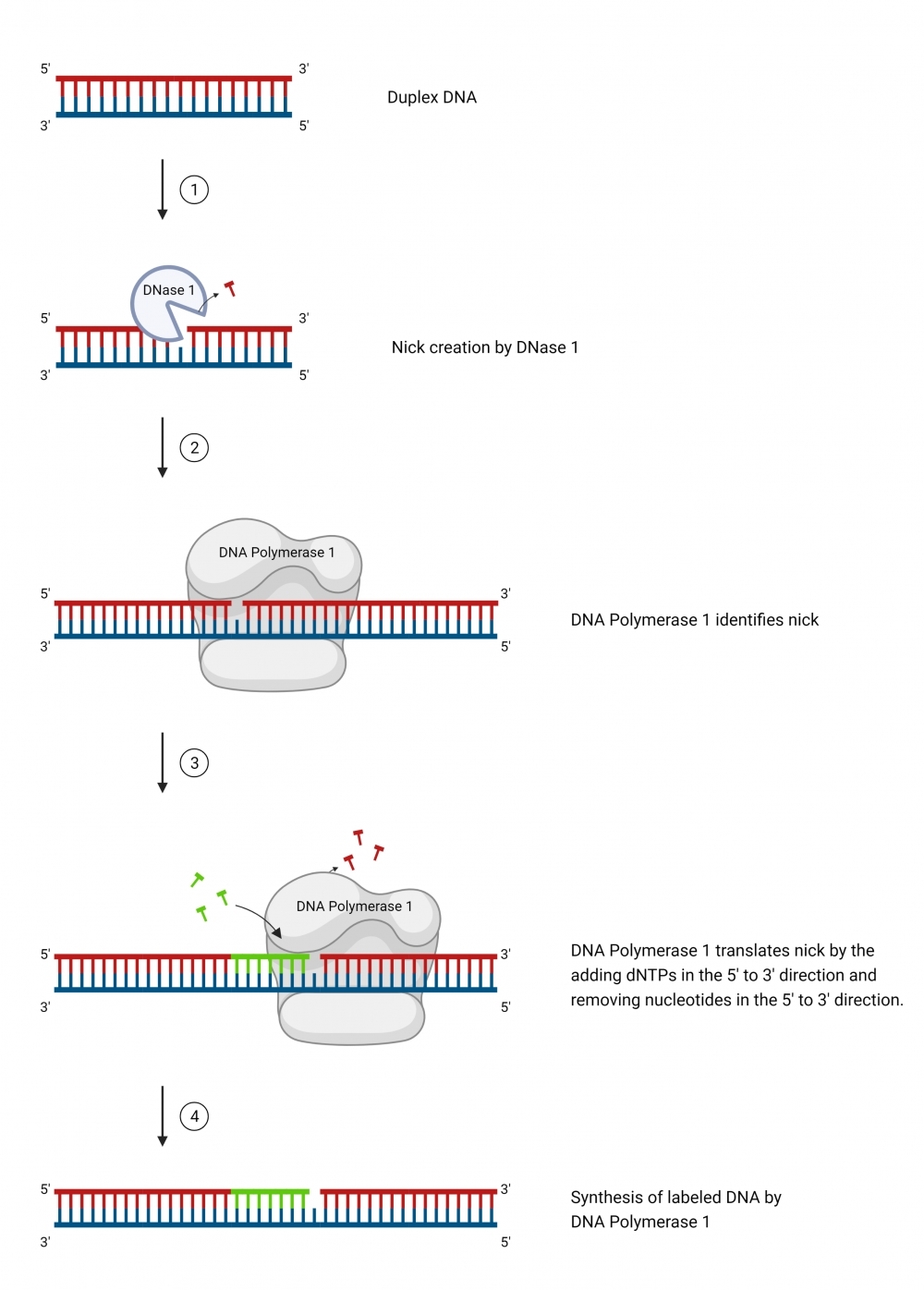
ReadiLink™ iFluor® 647 Nick Translation dsDNA Labelling Kit provides a simple and efficient way to label a double stranded DNA sample with the bright and photostable iFluor® 647 dye. The labelling kit provides all necessary reagents for a complete workflow required for DNA labelling. This method utilizes a combination of DNAse and DNA polymerase to nick one strand of the DNA helix, to which iFluor® 647 dye is conjugated. In addition, the kit allows the user to optimize incorporation and product size by adjusting the ratio of iFluor® 647-dUTP conjugate to dTTP. It is compatible with a wide variety of sample materials, including bacterial artificial chromosome (BAC) DNA, human genomic DNA, purified PCR products, supercoiled and linearized plasmid DNA. The resulted iFluor® 647-labeled DNAs can be used in a variety of molecular biology techniques such as fluorescence in situ hybridization (FISH).

| Catalog | Size | Price | Quantity |
|---|---|---|---|
| 17404 | 10 Reactions | Price | |
| 17405 | 20 Reactions | Price |
| Correction factor (260 nm) | 0.03 |
| Correction factor (280 nm) | 0.03 |
| Correction factor (656 nm) | 0.0793 |
| Extinction coefficient (cm -1 M -1) | 250000 1 |
| Excitation (nm) | 656 |
| Emission (nm) | 670 |
| Quantum yield | 0.25 1 |
| H-phrase | H303, H313, H333 |
| Hazard symbol | XN |
| Intended use | Research Use Only (RUO) |
| R-phrase | R20, R21, R22 |
| UNSPSC | 12171501 |
| Other instruments | Thermal Cycler |
| Telephone | |
| Fax | |
| sales@aatbio.com | |
| International | See distributors |
| Bulk request | Inquire |
| Custom size | Inquire |
| Technical Support | Contact us |
| Request quotation | Request |
| Purchase order | Send to sales@aatbio.com |
| Shipping | Standard overnight for United States, inquire for international |

