TAQuest™ qPCR Master Mix for TaqMan Probes
No ROX
TAQuest™ qPCR Master Mix for TaqMan Probes is a ready-to-use 2X solution optimized for qPCR and 2-step RT-qPCR compatible with TaqMan gene expression assays. The master mix provides all of the essential components including our proprietary TAQuest™ Hot Start Taq DNA Polymerase enzyme and dNTPs in an optimized PCR buffer, except the template, primers and probes. The Hot Start Taq DNA polymerase allows you to set up a PCR reaction at room temperature, thus minimizing non-specific product formation. The optimized composition ensures PCR specificity and sensitivity with all sample types such as genomic, plasmid, viral and cDNA templates. TAQuest™ qPCR Master Mix for TaqMan Probes has been designed to be used for duplex reactions using internal positive controls. This master mix does not contain a ROX reference dye.
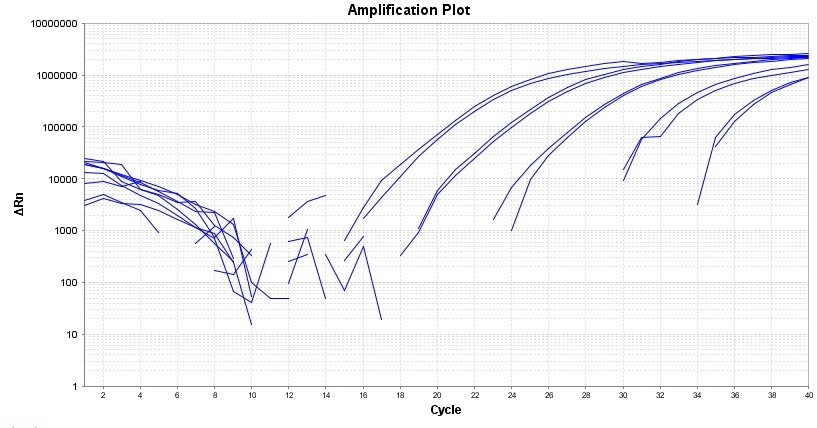

| Catalog | Size | Price | Quantity |
|---|---|---|---|
| 17282 | 1 mL | Price | |
| 17283 | 5 mL | Price |
Physical properties
| Solvent | Water |
Storage, safety and handling
| H-phrase | H303, H313, H333 |
| Hazard symbol | XN |
| Intended use | Research Use Only (RUO) |
| R-phrase | R20, R21, R22 |
| Storage | Freeze (< -15 °C); Minimize light exposure |
| UNSPSC | 12171501 |
Instrument settings
| qPCR | |
| Instrument specification(s) | Filter based on probes |
Contact us
| Telephone | |
| Fax | |
| sales@aatbio.com | |
| International | See distributors |
| Bulk request | Inquire |
| Custom size | Inquire |
| Technical Support | Contact us |
| Request quotation | Request |
| Purchase order | Send to sales@aatbio.com |
| Shipping | Standard overnight for United States, inquire for international |
Page updated on April 16, 2026
