Amplite® Colorimetric Aldehyde Quantitation Kit
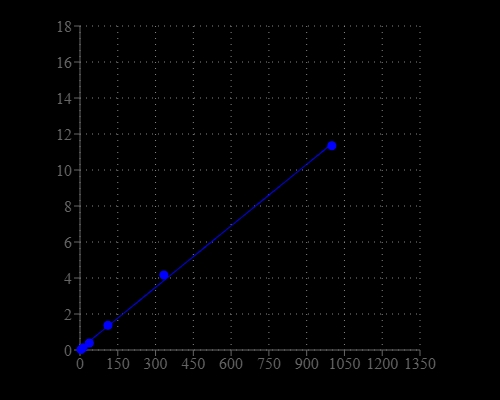
Rapid and accurate measurement of aldehydes is important task for biological research, food industry, chemical research and environmental pollution surveillance. There are few reagents or assay kits available for quantifying the amount of aldehydes. Most of existing aldehyde test methods is based on separations either by the tedious and expensive HPLC-MS or GC-MS. Our Amplite® Colorimetric Aldehyde Quantitation kit uses a proprietary dye that generates a chromogenic product upon reacting with an aldehyde. The kit provides a sensitive, one-step colorimetric method to detect as little as 1 nanomole of aldehyde in a 100 µL assay volume (10 µM; Figure 1). The assay can be performed in a convenient 96-well or 384-well microtiter-plate format and easily adapted to automation with no separation steps required. Its signal can be easily read by absorbance microplate reader at 550 nm.

| Catalog | Size | Price | Quantity |
|---|---|---|---|
| 10051 | 200 Tests | Price |
Storage, safety and handling
| H-phrase | H303, H313, H333 |
| Hazard symbol | XN |
| Intended use | Research Use Only (RUO) |
| R-phrase | R20, R21, R22 |
| UNSPSC | 12171501 |
Instrument settings
| Absorbance microplate reader | |
| Absorbance | 550 nm |
| Recommended plate | Clear bottom |
Contact us
| Telephone | |
| Fax | |
| sales@aatbio.com | |
| International | See distributors |
| Bulk request | Inquire |
| Custom size | Inquire |
| Technical Support | Contact us |
| Request quotation | Request |
| Purchase order | Send to sales@aatbio.com |
| Shipping | Standard overnight for United States, inquire for international |
Page updated on June 5, 2026
