Protonex™ Green 500, SE
This product has been discontinued. It is replaced by Catalog Number 21219, which has improved water solubility.
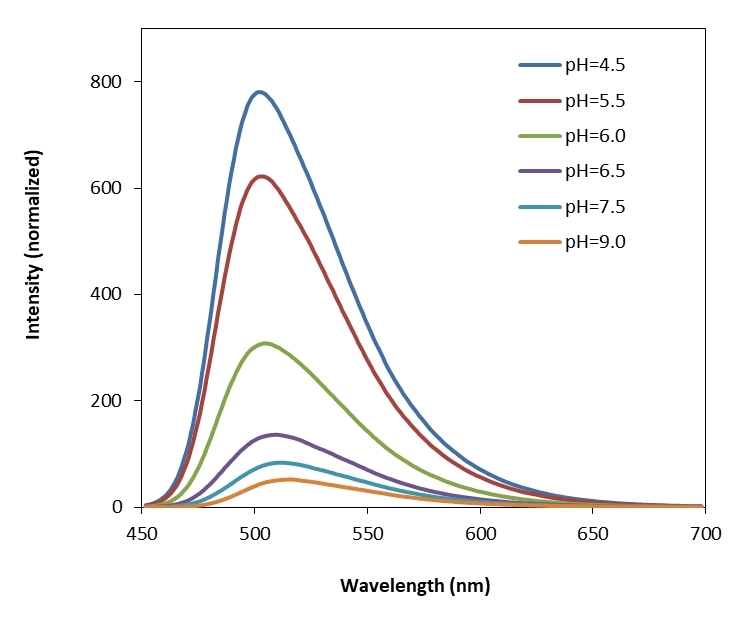
Protonex™ Green dye demonstrated pH-dependent fluorescence. Unlike most of the existing fluorescent dyes that are more fluorescent at higher pH, acidic conditions enhance the fluorescence of Protonex™ Green dye. The fluorescence of Protonex™ Green dye increases as pH decreases from neutral to the acidic. The lack of fluorescence outside the cell eliminates the wash steps. Protonex™ Green dye provides a powerful tool to monitor acidic cell compartments such as endosomes and lysosomes. Protonex™ Green dye is non-fluorescent outside the cells, but fluoresces brightly green in acidic compartments (such as phagosomes, lysosomes and endosomes). This Protonex™ Green enables the specific detection of cellular acidic compartments with reduced signal variability and improved accuracy for imaging or flow applications. Protonex™ Green has the spectral properties similar to those of FITC, making the common filter set of FITC readily available to the assays of Protonex™ Green. This amino-reactive Protonex™ Green derivative can be used for preparing Protonex™ Green conjugates as fluorescent pH probes.

| Catalog | Size | Price | Quantity |
|---|---|---|---|
| 21216 | 1 mg | Price |
Physical properties
| Molecular weight | 539.54 |
| Solvent | DMSO |
Spectral properties
| Extinction coefficient (cm -1 M -1) | 4000 |
| Excitation (nm) | 445 |
| Emission (nm) | 503 |
Storage, safety and handling
| H-phrase | H303, H313, H333 |
| Hazard symbol | XN |
| Intended use | Research Use Only (RUO) |
| R-phrase | R20, R21, R22 |
| Storage | Freeze (< -15 °C); Minimize light exposure |
| UNSPSC | 12352200 |
Contact us
| Telephone | |
| Fax | |
| sales@aatbio.com | |
| International | See distributors |
| Bulk request | Inquire |
| Custom size | Inquire |
| Technical Support | Contact us |
| Request quotation | Request |
| Purchase order | Send to sales@aatbio.com |
| Shipping | Standard overnight for United States, inquire for international |
Page updated on June 9, 2026

