Protonex™ Red 670-Latex Bead Conjugate
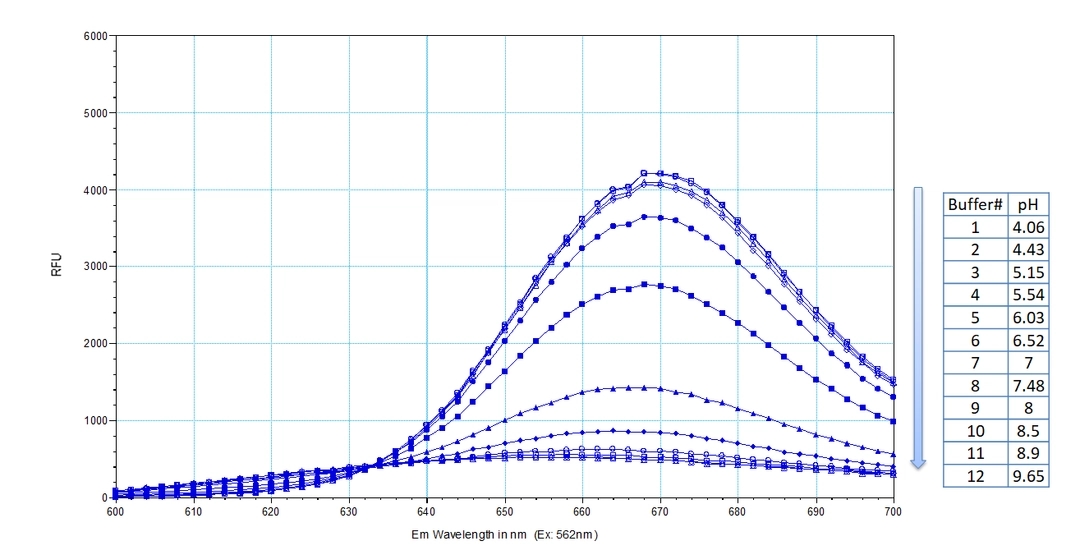
Protonex™ Red 670-Latex Bead Conjugates exhibit unique pH-dependent fluorescence properties. Unlike most existing fluorescent dyes that typically increase in fluorescence at higher pH levels, Protonex™ Red 670-Latex Bead Conjugates become significantly more fluorescent under acidic conditions. This characteristic makes them an ideal tool for studying phagocytosis and its modulation by drugs or environmental factors. The beads display minimal fluorescence outside of cells, eliminating the need for wash steps and enhancing their utility for live-cell imaging. Protonex™ Red 670-Latex Bead Conjugates are particularly effective in highlighting acidic cellular compartments such as phagosomes, lysosomes, and endosomes, where they emit a bright red fluorescence. Additionally, Protonex™ Red 670-Latex Bead Conjugates can be used in combination with green fluorescent dyes like GFP, Fluo-8®, calcein, or FITC-labeled antibodies for multiplexed cell functional analysis. Their spectral properties are similar to those of Cy5, allowing for the use of common Cy5 filter sets in assays involving Protonex™ Red 670, further facilitating their integration into existing experimental setups.

| Catalog | Size | Price | Quantity |
|---|---|---|---|
| 21222 | 1 mL | Price |
Spectral properties
| Excitation (nm) | 643 |
| Emission (nm) | 660 |
Storage, safety and handling
| H-phrase | H303, H313, H333 |
| Hazard symbol | XN |
| Intended use | Research Use Only (RUO) |
| R-phrase | R20, R21, R22 |
| Storage | Refrigerated (2-8 °C); Minimize light exposure |
| UNSPSC | 12352200 |
Instrument settings
| Fluorescence microscope | |
| Excitation | Cy5 filter set |
| Emission | Cy5 filter set |
| Recommended plate | Black wall/clear bottom |
Contact us
| Telephone | |
| Fax | |
| sales@aatbio.com | |
| International | See distributors |
| Bulk request | Inquire |
| Custom size | Inquire |
| Technical Support | Contact us |
| Request quotation | Request |
| Purchase order | Send to sales@aatbio.com |
| Shipping | Standard overnight for United States, inquire for international |
Page updated on June 5, 2026

