Live or Dead™ Cell Viability Assay Kit
Red/Blue Dual Fluorescence
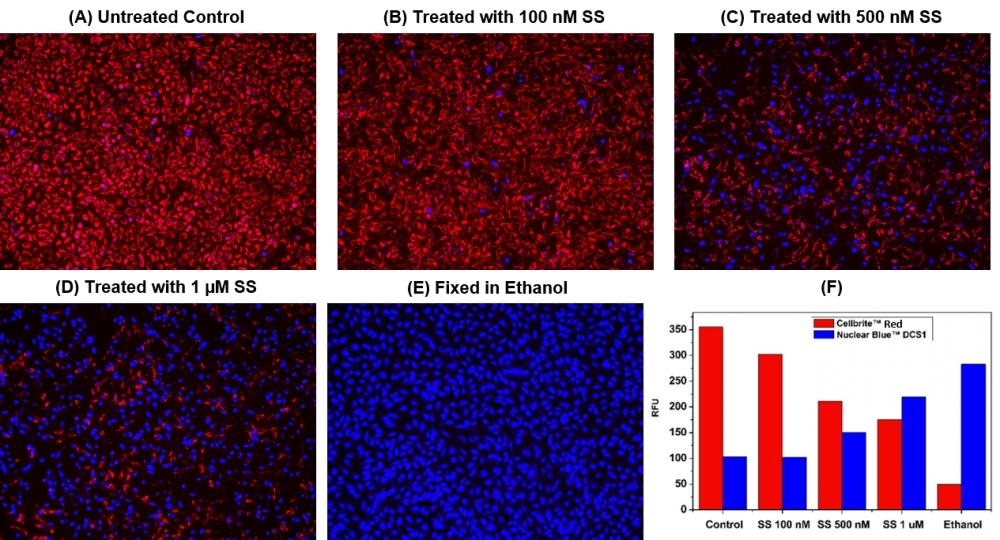
This Live or Dead™ Cell Viability Assay Kit uses two fluorescent indicators: Cellbrite™ Red (Ex/Em = 613/631 nm) for labeling viable cells and a cell-impermeable DNA-binding dye Nuclear Blue™ DCS1 (Ex/Em = 360/450 nm) for labeling dead cells with damaged membranes. Cells grown in black-wall plates can be stained and quantified in less than two hours. The assay is more robust and accurate than the other viability assays. It can be readily adapted for a wide variety of fluorescence platforms such as microplate assays, fluorescence microscopes, and flow cytometry. The kit provides all the essential components with an optimized assay protocol. It is suitable for both proliferating and non-proliferating cells (either suspension or adherent cells).

| Catalog | Size | Price | Quantity |
|---|---|---|---|
| 22788 | 200 Tests | Price |
Spectral properties
| Excitation (nm) | 612 |
| Emission (nm) | 630 |
Storage, safety and handling
| H-phrase | H303, H313, H333 |
| Hazard symbol | XN |
| Intended use | Research Use Only (RUO) |
| R-phrase | R20, R21, R22 |
| UNSPSC | 12352200 |
Instrument settings
| Fluorescence microscope | |
| Excitation | Cy5 filter (alive), DAPI filter (dead) |
| Emission | Cy5 filter (alive), DAPI filter (dead) |
| Recommended plate | Black wall/clear bottom |
| Fluorescence microplate reader | |
| Excitation | 610, 360 nm |
| Emission | 650, 450 nm |
| Cutoff | 630, 420 nm |
| Recommended plate | Black wall/clear bottom |
| Instrument specification(s) | Bottom read mode |
Contact us
| Telephone | |
| Fax | |
| sales@aatbio.com | |
| International | See distributors |
| Bulk request | Inquire |
| Custom size | Inquire |
| Technical Support | Contact us |
| Request quotation | Request |
| Purchase order | Send to sales@aatbio.com |
| Shipping | Standard overnight for United States, inquire for international |
Page updated on August 5, 2024

