Portelite™ Fluorimetric Protein Quantitation Kit
Optimized for CytoCite™ and Qubit™ Fluorometers
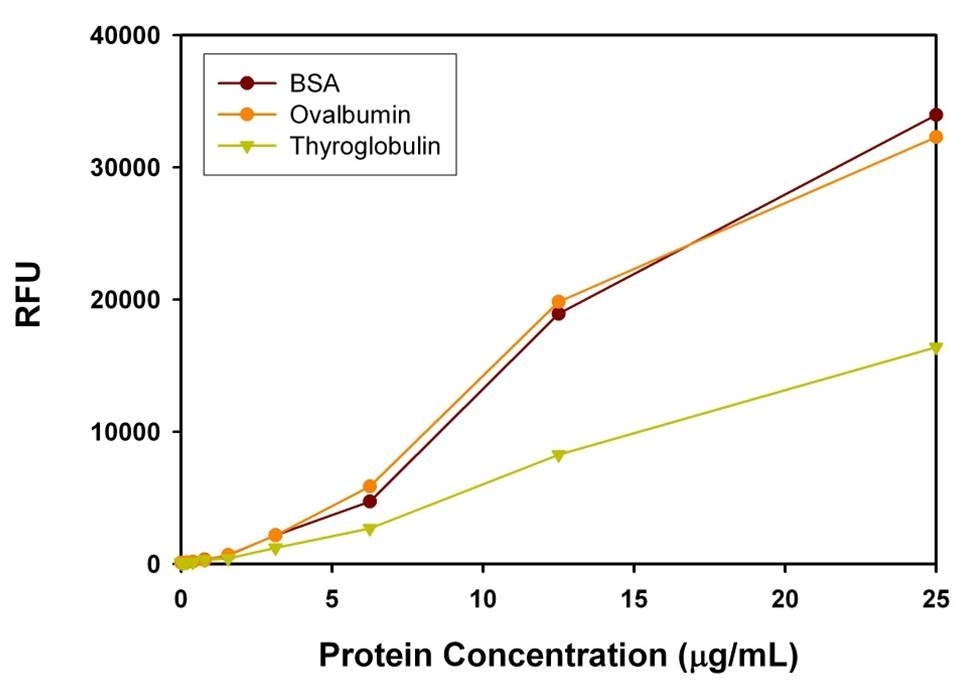
Protein quantification is an essential task in protein purification, electrophoresis, cell biology, molecular biology and other research applications. Biuret, Lowry, BCA and Bradford assays are routinely used for estimating protein concentration. However, these colorimetric assays are less sensitive, and require large sample volume to ensure accuracy. Our Portelite™ Fluorimetric Protein Quantitation Kit is significantly more sensitive than existing colorimetric protein measurements, e.g., Bradford and Bicinchoninic acid (BCA) assays. Prolite™ Orange used in the kit is non-fluorescent in aqueous solution, but reacts rapidly with proteins and generates bright fluorescence. The Portelite™ Fluorimetric Protein Quantitation Kit provides a simple method for quantifying protein concentration in solutions. The assay has dynamic range from 12.5 ug/mL to 5 mg/mL of BSA. The kit is optimized for Cytocite™ and Qubit™ fluorometers. It can be used for (1) studying protein/protein interactions; (2) measuring column fractions after affinity chromatography; (3) estimating recovery of membrane proteins from cell extract; and (4) high-throughput screening of fusion proteins.

| Catalog | Size | Price | Quantity |
|---|---|---|---|
| 11109 | 100 Tests | Price | |
| 11111 | 500 Tests | Price |
Storage, safety and handling
| H-phrase | H303, H313, H333 |
| Hazard symbol | XN |
| Intended use | Research Use Only (RUO) |
| R-phrase | R20, R21, R22 |
| UNSPSC | 12171501 |
Instrument settings
| Qubit Fluorometer | |
| Excitation | 480 nm |
| Emission | 510-580 nm |
| Instrument specification(s) | 0.2 mL, thin-wall PCR tube |
| CytoCite Fluorometer | |
| Excitation | 480 nm |
| Emission | 510-580 nm |
| Instrument specification(s) | 0.2 mL, thin-wall PCR tube |
Contact us
| Telephone | |
| Fax | |
| sales@aatbio.com | |
| International | See distributors |
| Bulk request | Inquire |
| Custom size | Inquire |
| Technical Support | Contact us |
| Request quotation | Request |
| Purchase order | Send to sales@aatbio.com |
| Shipping | Standard overnight for United States, inquire for international |
Page updated on April 30, 2026
