RatioWorks™ PDMPO Dextran
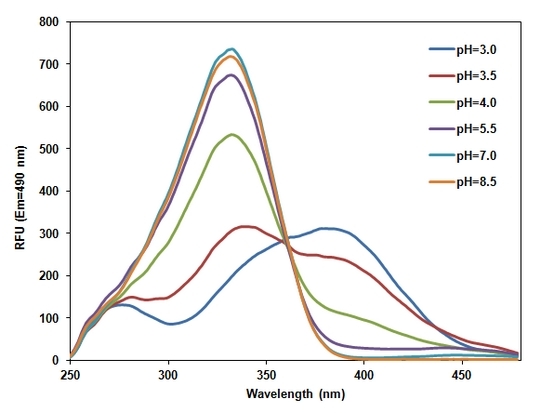
The existing pH probes are ill-adapted to study acidic organelles such as lysosomes, endosomes, phagosomes, spermatozoa and acrosomes because their fluorescence is significantly reduced at lower pH. The growing potential of ratio imaging is significantly limited by the lack of appropriate fluorescent probes for acidic organelles although ratio imaging has received intensive attention in the past few decades. RatioWorks™ PDMPO is characterized as acidotropic dual-excitation and dual-emission pH probe. It emits intense yellow fluorescence at lower pH and gives intense blue fluorescence at higher pH. This unique pH-dependent fluorescence makes RatioWorks™ PDMPO an ideal pH probe for acidic organelles with pKa = 4.47. Additionally, the very large Stokes shift and excellent photostability of RatioWorks™ PDMPO make it an excellent fluorescent acidotropic reagent for fluorescence imaing and flow cytometry applications. The unique fluorescence properties of RatioWorks™ PDMPO might give researchers a new tool with which to study endocytosis, phagocytosis and acidic organelles of live cells. RatioWorks™ PDMPO can be well excited by the violet laser at 405 nm for flow cytometric applications. This RatioWorks™ PDMPO SE can be readily used to make a variety of bioconjugates for imaging or flow applications, enabling the specific detection of phagocytosis and endocytosis with reduced signal variability and improved accuracy. These conjugates can be also used for multiplexing cell functional analysis with green dyes such as GFP, Fluo-8®, calcein, or FITC-labeled antibodies. The short emission band of RatioWorks™ PDMPO is ~450 nm while the longer emission is ~550 nm, making the common filter sets of Pacific Blue and Pacific Orange readily available to the assays of RatioWorks™ PDMPO.

| Catalog | Size | Price | Quantity |
|---|---|---|---|
| 21211 | 1 mg | Price |
Physical properties
| Molecular weight | ~10000 |
| Solvent | Water |
Spectral properties
| Excitation (nm) | 333 |
| Emission (nm) | 531 |
Storage, safety and handling
| H-phrase | H303, H313, H333 |
| Hazard symbol | XN |
| Intended use | Research Use Only (RUO) |
| R-phrase | R20, R21, R22 |
| Storage | Freeze (< -15 °C); Minimize light exposure |
| UNSPSC | 12352200 |
Instrument settings
| Fluorescence microplate reader | |
| Excitation | 360 nm |
| Emission | 450, 540 nm |
| Cutoff | 420, 475 nm |
| Recommended plate | Black wall/Clear bottom |
| Instrument specification(s) | Bottom read mode |
Contact us
| Telephone | |
| Fax | |
| sales@aatbio.com | |
| International | See distributors |
| Bulk request | Inquire |
| Custom size | Inquire |
| Technical Support | Contact us |
| Request quotation | Request |
| Purchase order | Send to sales@aatbio.com |
| Shipping | Standard overnight for United States, inquire for international |
Page updated on June 6, 2026

