Cell Meter™ Intracellular Fluorimetric Hydrogen Peroxide Assay Kit
Green Fluorescence
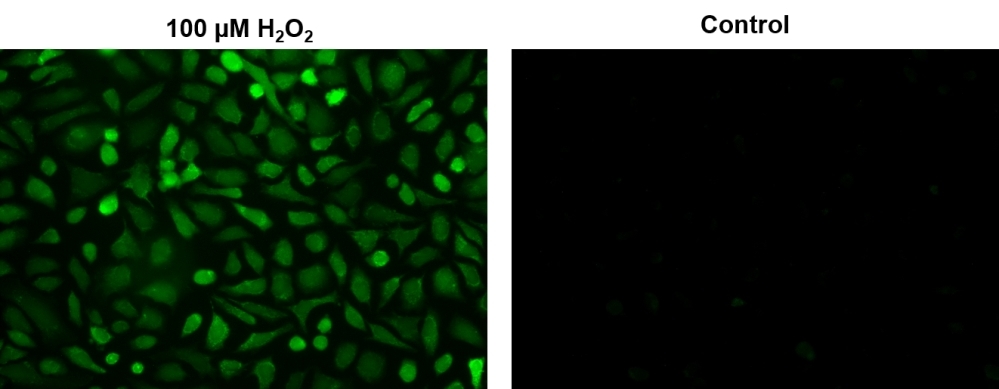
Hydrogen peroxide (H2O2) is a reactive oxygen metabolic by-product that serves as a key regulator for a number of oxidative stress-related states. It is involved in a number of biological events that have been linked to asthma, atherosclerosis, diabetic vasculopathy, osteoporosis, a number of neurodegenerative diseases and Down's syndrome. Perhaps the most intriguing aspect of hydrogen peroxide biology is the recent report that antibodies have the capacity to convert molecular oxygen into hydrogen peroxide to contribute to the normal recognition and destruction processes of the immune system. Measurement of this reactive species will help to determine how oxidative stress modulates varied intracellular pathways. This Cell Meter™ Hydrogen Peroxide Assay Kit uses our unique OxiVision™ Green hydrogen peroxide sensor to quantify hydrogen peroxide in live cells. OxiVision™ Green is cell-permeable, and generates the green fluorescence when it reacts with hydrogen peroxide. The kit is an optimized 'mix and read' assay format that is compatible with HTS liquid handling instruments.

| Catalog | Size | Price | Quantity |
|---|---|---|---|
| 11503 | 200 Tests | Price |
Spectral properties
| Excitation (nm) | 498 |
| Emission (nm) | 517 |
Storage, safety and handling
| H-phrase | H303, H313, H333 |
| Hazard symbol | XN |
| Intended use | Research Use Only (RUO) |
| R-phrase | R20, R21, R22 |
| UNSPSC | 12171501 |
Instrument settings
| Fluorescence microscope | |
| Excitation | FITC channel |
| Emission | FITC channel |
| Recommended plate | Black wall/clear bottom |
| Fluorescence microplate reader | |
| Excitation | 490 nm |
| Emission | 525 nm |
| Cutoff | 515 nm |
| Recommended plate | Black wall/clear bottom |
| Instrument specification(s) | Bottom read mode |
Contact us
| Telephone | |
| Fax | |
| sales@aatbio.com | |
| International | See distributors |
| Bulk request | Inquire |
| Custom size | Inquire |
| Technical Support | Contact us |
| Request quotation | Request |
| Purchase order | Send to sales@aatbio.com |
| Shipping | Standard overnight for United States, inquire for international |
Page updated on May 11, 2026

