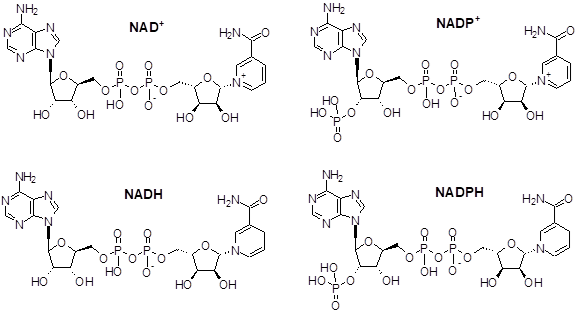
NAD/NADH and NADP/NADPH
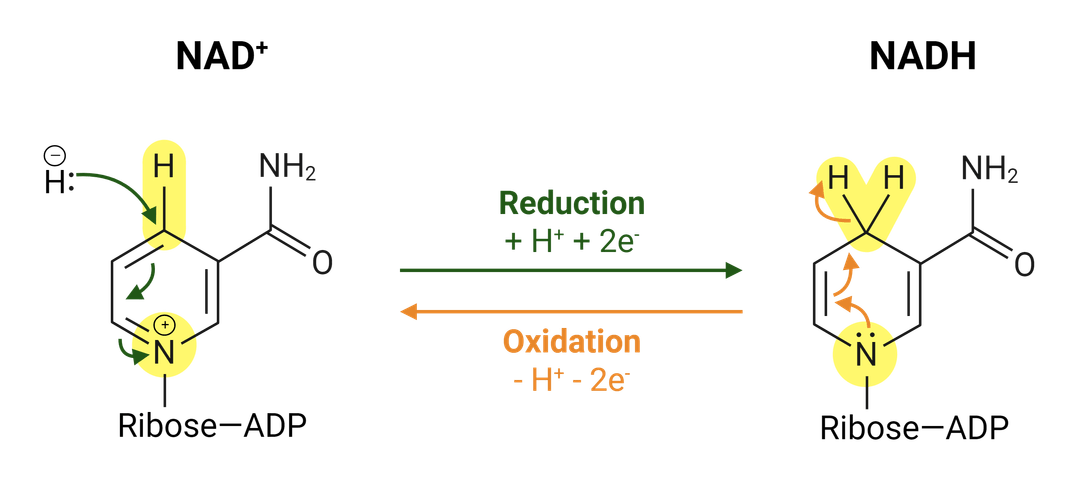
Nicotinamide adenine dinucleotide (NAD+) and NAD+ phosphatase (NADP+) are coupled redox cofactors associated with maintaining chemical equilibrium in the cellular environment with active roles in many signal pathways, including cellular metabolism. NAD(P)+ are critical in managing oxidative and reducing stress, and are classified under coenzymes in the oxidoreductase family. NAD+ reduces to NADH through dehydrogenases, and phosphorylates to NADP+ through associated kinases, where each play a role in glycolysis and mitochondrial function. NAD+ is synthesized through various methods in the cytosol, nucleus and mitochondria, though reactions are highly compartmentalized within the microbiome due to limited bioavailability of precursors and partnered enzymes.
Table of Contents
Pathways of NAD(P)+
De novo Synthesis
Tryptophan (Trp) can be converted to NAD+ by the kynurenine pathway (KP) which is activated by one of two dioxygenase molecules, abbreviated TDO or IDO. Hydroxylated kynurenine undergoes three more enzymatic conformations before becoming its final form; first firming an unstable amine (ACMS), then Quinolinic Acid (QA), next nicotinic acid (NA) mononucleotide (NAMN), and finally to NA adenine dinucleotide (NAAD), before becoming NAD+. The final conversion of NAAD to NAD+ is mediated by ATP-dependent NAD+ synthetases (NADSYNs) which are found readily in the liver, kidney, and small intestine.
| Assaywise Letters: | Application Notes: |
The Preiss-Handler and Salvage Pathways
The Priess-Handler pathway is a three step process where NAD+ is produced with the addition of ATP and glutamine as a nitrogen donor, involving NAMN and NAAD as intermediate metabolites. This pathway requires NA as starting material, commonly known as the vitamin niacin, which is the limiting substrate. The salvage pathway creates NAM mononucleotide (NMN) from either Nicotinamide (NAM) or NAM riboside (NR) along with the aid of ATP. NMN adenylyltransferases (NMNATs) then catalyzes the conversion of this intermediate into NAD+, also involved in the De novo synthesis platform, that has three associated isoforms, each with distinct localizations in the cell.
| Application Notes: |
NADP+ Biosynthesis

Chemical structures of oxidized and reduced NADP(H) as associated with the ReadiUse™ NADP Regenerating Kit.
Table 1. Biochemical assays for measuring ATP activity, formation or depletion.
| Assay ▲ ▼ | Ex/Abs (nm) ▲ ▼ | Em (nm) ▲ ▼ | Cutoff (nm)¹ ▲ ▼ | Microplate Type ▲ ▼ | Unit Size ▲ ▼ | Cat No. ▲ ▼ |
| PhosphoWorks™ Fluorimetric ATP Assay Kit | 540 | 590 | 570 | Solid black | 100 tests | 21620 |
| PhosphoWorks™ Colorimetric ATP Assay Kit | 570 | - | - | Clear bottom | 100 tests | 21617 |
| PhosphoWorks™ Luminometric ATP Assay Kit *Maximized Luminescence* | - | - | - | Solid white | 1 plate | 21610 |
| PhosphoWorks™ Luminometric ATP Assay Kit *Maximized Luminescence* | - | - | - | Solid white | 10 plate | 21621 |
| Cell Meter™ Live Cell ATP Assay Kit | - | - | - | Black wall/clear bottom | 100 Tests | 23015 |
NAD(P)+ Regulation

Chemical structure for ADP-ribose-pNP, a colorimetric substrate for assessing activity of poly(ADP-ribose)polymerase (PARP) enzymes.
| Assaywise Letters: | Datasets: |
NAD(P)+ Physiology
NAD(P)+ pathways are also inter connected in a number of other signaling pathways. The conversion of NAD+ to NADH stimulates glycolysis, fatty acid oxidation and the tricarboxylic acid cycle. NADH altering back to NAD+ affects lactic acid degeneration, PUFA desaturation, and the mitochondrial electron transport chain. NADP+ conversion to NADPH vitalizes the pentose phosphate pathway, which when transformed back into the monovalent cations NADP+ is involved in antioxidant defense, anabolic reactions and NOX signaling.
NAD+ is also involved in RNA capping in many species, and aids in DNA ligation and repair through adenylation. Besides simply maintaining redox homeostasis, NAD(P)+ also donate electrons in reactive oxygen species (ROS) generation and play a part in the creation of antioxidant species. Due to this role, NAD(P)+ is associated with maintaining genomic equanimity and gene expression manipulation. It is apparent that NAD(P)+ is networked to many different signal pathways, issues in NAD(P)+ metabolism could lead to a multitude of potential negative ensuing effects associated with immune and inflammatory disease.
Measuring NAD(P)+/NAD(P)H
Biochemical Techniques
High-performance liquid chromatography (HPLC) can be used to detect and quantify metabolites produced from NAD(P)+, and is usually coupled with mass spectrometry (LC/MS). The idea is that NAD(P)+ biosynthesis, which positively supports body function, can be monitored through NAD(P)+ intermediate production, namely NMN and NR. Intermediates are identified through acid extraction and NAD(P)+ concentration can be obtained. Enzymatic cycling assays, capillary electrophoresis, and isotope-labeling techniques have also been used, usually alongside LC/MS, though these processes measure total, cumulative NAD(P)+ and do not distinguish between subcomponent concentrations. Each method can also quantify the reduced coenzyme forms, with reproducible and rapid results through fragmentation.
Fluorescence
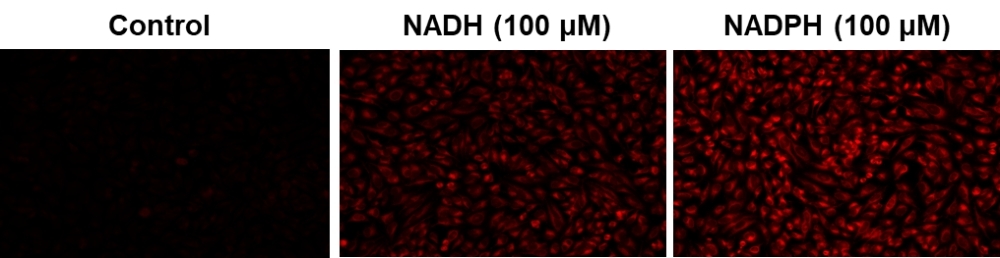
Fluorescence images of NADH/NADPH in HeLa cells using Cell Meter™ Intracellular NADH/NADPH Fluorescence Imaging Kit.
Genetically encoded fluorescent sensors are also utilized to detect NAD(P)+/H changes in real-time, and techniques also target specific redox reactions, thiols, and ROS. These techniques can also be applied in a high-throughput setting, using many available equipment types like microplate readers, flow cytometry, fluorescence microscopy and other optical imaging systems. Benefits to said biosensors are acute fluorescence, rapid excitation, and subcellular organelle targeting where cytosolic, compartmentalized and mitochondrial NAD(P)+ pools can be separately quantified. Fluorescent techniques still have a long way to go, and easily distinguishing between NADH and NADPH levels in samples remains an issue. Certain biosensors are also not totally impassive to alterations in pH, so consideration must be taken in this regard as well.
P-Magnetic Resonance Spectroscopy (P-MRS)
P-MRS techniques have been used to measure NAD+/H levels in vivo by employing an ultra-high magnetic field, specifically in areas of the brain. Results can be interpreted through analysis by charts showing chemical shifts between the resonance frequency of the observed proton and hydrogens on tetramethylsilane. Through this method, ATP and NAD+/H pools can be distinguished in a minimally invasive manner to living tissue. Therapeutic advances so far have been limited to the low-resolution quality provided by low magnetic fields needed for clinical applications; however progress continues to be made on this forefront.
Product Ordering Information
Table 2. NAD/NADH & NADP/NADPH Assay Comparison
Table 3. Ordering Info for NAD/NADH Products
| Cat# ▲ ▼ | Product Name ▲ ▼ | Unit Size ▲ ▼ |
| 15257 | Amplite® Fluorimetric Total NAD and NADH Assay Kit *Red Fluorescence* | 400 Tests |
| 15258 | Amplite® Colorimetric Total NAD and NADH Assay Kit | 400 Tests |
| 15261 | Amplite® Fluorimetric NADH Assay Kit *Red Fluorescence* | 400 Tests |
| 15263 | Amplite® Fluorimetric NAD/NADH Ratio Assay Kit *Red Fluorescence* | 250 Tests |
| 15271 | Amplite® Colorimetric NADH Assay Kit | 400 Tests |
| 15273 | Amplite® Colorimetric NAD/NADH Ratio Assay Kit | 250 Tests |
| 15275 | Amplite® Colorimetric Total NAD and NADH Assay Kit *Enhanced Sensitivity* | 400 Tests |
| 15280 | Amplite® Fluorimetric NAD Assay Kit *Blue Fluorescence* | 200 Tests |
Table 4. Ordering Info for NADP/NADPH Products
| Cat# ▲ ▼ | Product Name ▲ ▼ | Unit Size ▲ ▼ |
| 15260 | Amplite® Colorimetric Total NADP and NADPH Assay Kit | 400 Tests |
| 15262 | Amplite® Fluorimetric NADPH Assay Kit *Red Fluorescence* | 400 Tests |
| 15264 | Amplite® Fluorimetric NADP/NADPH Ratio Assay Kit *Red Fluorescence* | 250 Tests |
| 15265 | ReadiUse™ NADPH Regenerating Kit | 1000 Tests |
| 15272 | Amplite® Colorimetric NADPH Assay Kit | 400 Tests |
| 15274 | Amplite® Colorimetric NADP/NADPH Ratio Assay Kit | 250 Tests |
| 15276 | Amplite® Colorimetric Total NADP and NADPH Assay Kit *Enhanced Sensitivity* | 400 Tests |
| 15281 | Amplite® Fluorimetric NADP Assay Kit *Blue Fluorescence* | 200 Tests |
| 15295 | Cell Meter™ Intracellular NADH/NADPH Fluorescence Imaging Kit *Deep Red Fluorescence* | 100 Tests |
References
Xiao, Wusheng, et al. “NAD(H) and NADP(H) Redox Couples and Cellular Energy Metabolism.” Antioxidants & Redox Signaling, vol. 28, no. 3, 2018, pp. 251-272., https://doi.org/10.1089/ars.2017.7216.
Xie, Na, et al. “NAD+ Metabolism: Pathophysiologic Mechanisms and Therapeutic Potential.” Signal Transduction and Targeted Therapy, vol. 5, no. 1, 2020, https://doi.org/10.1038/s41392-020-00311-7.
Kelly, Gregory. “How Is NAD+ Made? Preiss-Handler Pathway.” Neurohacker Collective, 30 Aug. 2019, https://neurohacker.com/how-is-nad-made-preiss-handler-pathway.
Yoshino, Jun, and Shin-ichiro Imai. “Accurate Measurement of Nicotinamide Adenine Dinucleotide (NAD+) with High-Performance Liquid Chromatography.” Sirtuins, 26 Feb. 2014, pp. 203-215., https://doi.org/10.1007/978-1-62703-637-5_14.
Zou, Yejun, et al. “Analysis of Redox Landscapes and Dynamics in Living Cells and in Vivo Using Genetically Encoded Fluorescent Sensors.” Nature Protocols, vol. 13, no. 10, 2018, pp. 2362-2386., https://doi.org/10.1038/s41596-018-0042-5.
Lu, Ming, et al. “In vivo31p MRS Assessment of Intracellular NAD Metabolites and NAD+/NADH Redox State in Human Brain at 4 T.” NMR in Biomedicine, vol. 29, no. 7, 2016, pp. 1010-1017., https://doi.org/10.1002/nbm.3559.